gdellipsoid
Small Python package to build an ellipsoidal cardiac geometry: tetrahedral mesh, endo/epicardial surfaces, fiber directions, and a Purkinje network.
What it does
- Geometry: Parametric ellipsoid shell discretized into a tetrahedral volume with endocardium (ENDO) and epicardium (EPI) labels.
- Fibers: Transmural fiber directions (default angles 80° / −80° endo/epi) via a Laplace-based rule, written as point/cell data.
- Purkinje: Fractal tree on the endocardial surface (
fractal_tree), saved as a line mesh. - Plot: Single view of volume mesh, surfaces, Purkinje tree, and fiber samples.
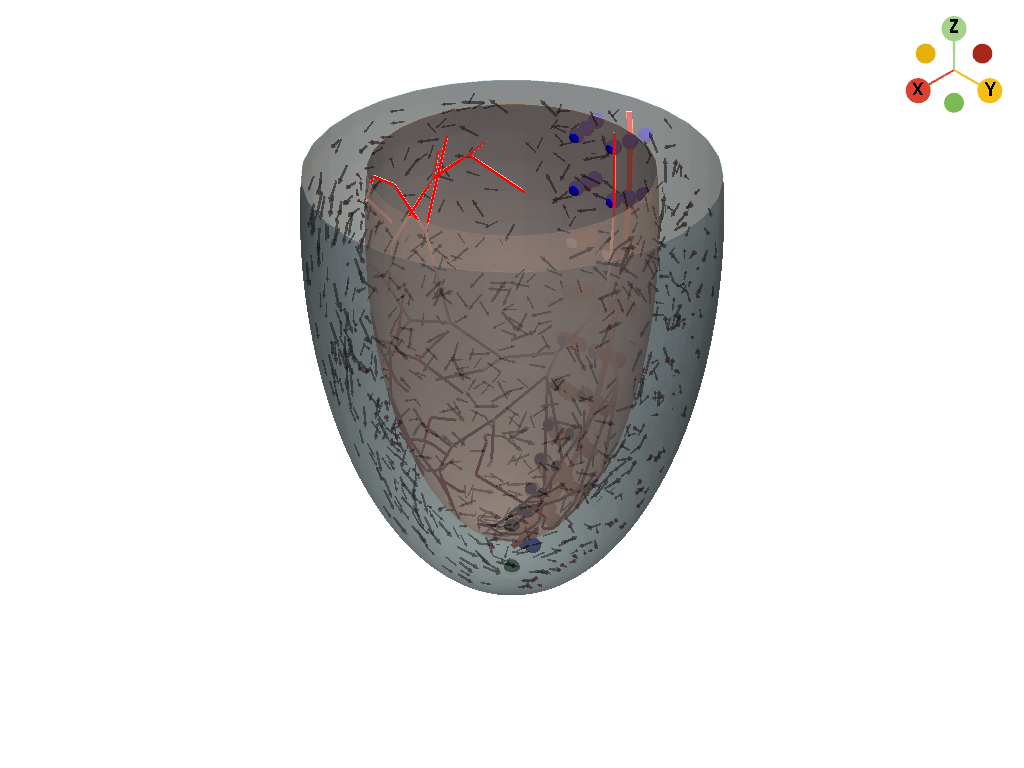
Output files
Outputs go to gdellipsoid/out/ by default: tetra.vtk, endo.vtk, epi.vtk, regular_surf.vtk, points.vtk, rings.obj, fiber meshes, purkinje_mesh.vtk, and a combined plot.
Main API
| Function / module | Role |
|---|---|
gdellipsoid.main.generate_ellipsoid(out) |
Build and save ellipsoid meshes (tetra, surfaces, points, rings). |
gdellipsoid.fibers.create_fibers(out, angles=(80, -80)) |
Compute and save fiber directions. |
gdellipsoid.purkinje.create_purkinje(out) |
Build Purkinje tree on endo and save mesh. |
gdellipsoid.plot.plot_all(out) |
Plot volume, surfaces, Purkinje, and fibers. |
Dependencies
Uses gdutils, gdmesh, pyvista, meshio, fractal_tree, numpy, scipy, and (for fibers) skfem.